- Clone
- A17035K (See other available formats)
- Regulatory Status
- RUO
- Other Names
- Beta-Secretase 1, Membrane-Associated Aspartic Protease 2, Beta-Site APP Cleaving Enzyme 1, Aspartyl Protease 2, Memapsin-2, BACE, ASP2, Transmembrane Aspartic Proteinase Asp2, Beta-Secretase 1 Precursor Variant 1, Beta-Site APP-Cleaving Enzyme 1
- Isotype
- Mouse IgG2b, κ
- Ave. Rating
- Submit a Review
- Product Citations
- publications

-
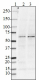
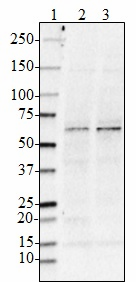
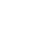
Western blot of purified anti-BACE1 antibody (clone A17035K). Lane 1: Molecular weight marker; Lane 2: 20 µg of total human brain lysate; Lane 3: 20 µg of human brain membrane lysate. The blot was incubated with 1.0 µg/mL of the primary antibody overnight at 4°C, followed by incubation with HRP-labeled goat anti-mouse IgG (Cat. No. 405301). Enhanced chemiluminescence (Cat. No. 426302) was used as the detection system. -
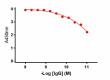
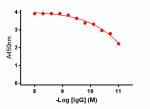
Direct ELISA of purified anti-BACE1 antibody (clone A17035K) binding to plate-immobilized recombinant human BACE1 (Cat. No. 719104). ELISA was performed by coating wells with 100ng of BACE1 protein. The wells were then incubated with the primary antibody at 37°C for 45 minutes, followed by incubation with horseradish peroxidase labeled goat anti-mouse secondary antibody. TMB (3, 3', 5, 5' tetramethylbenzidine, Cat. No. 421501) was used as the detection system.
| Cat # | Size | Price | Quantity Check Availability | Save | ||
|---|---|---|---|---|---|---|
| 852901 | 25 µg | $112 | ||||
| 852902 | 100 µg | $276 | ||||
Beta-site APP-cleaving enzyme 1 (BACE1), also known as aspartyl protease 2 (ASP-2) and memapsin 2, is a type I transmembrane aspartic protease with 501 amino acid that is related to the pepsin family and the retroviral aspartic proteases. The BACE1 catalytic domain contains aspartic protease active site motifs at amino acids 93-96 and 289-292, with each motif containing the highly conserved sequence defining aspartic proteases, D(T/S)G(T/S). BACE1 is expressed at low levels in most cell types of the body, although it is more highly expressed in neurons. In the brain, the protein is highly expressed in the substantia nigra, locus coruleus, and medulla oblongata. It is localized within acidic intracellular compartments including endosomes and trans-Golgi network (TGN). In the brain, BACE1 is involved in proteolytic processing of the amyloid precursor protein (APP) which generates the b amyloid peptide- a peptide that is associated with Alzheimer’s disease. BACE1 cleaves APP at the N-terminus of the b amyloid peptide sequence, between residues 671 and 672, which leads to extracellular release of b-cleaved soluble APP, and a corresponding cell-associated C-terminal fragment which is later released by g-secretase. The physiological function of BACE1 is not clearly understood, although it may modulate sodium currents by cleavage of voltage-gated Na+ channel accessory subunits (Navβ1-4). Also, BACE1 has been implicated in the regulation of myelination and myelin sheath thickness and in the control of motor coordination via cleavage of neuregulin-1 and neuregulin-3, and also in axon guidance via cleavage of close homolog of L1 (CHL1).
Product DetailsProduct Details
- Verified Reactivity
- Human
- Antibody Type
- Monoclonal
- Host Species
- Mouse
- Immunogen
- Recombinant human BACE1 protein.
- Formulation
- Phosphate-buffered solution, pH 7.2, containing 0.09% sodium azide.
- Preparation
- The antibody was purified by affinity chromatography.
- Concentration
- 0.5 mg/ml
- Storage & Handling
- The antibody solution should be stored undiluted between 2°C and 8°C.
- Application
-
WB - Quality tested
Direct ELISA - Verified - Recommended Usage
-
Each lot of this antibody is quality control tested by Western blotting. For Western blotting, the suggested use of this reagent is 1.0 - 5.0 µg per ml. For ELISA Capture applications, a concentration range of 0.002 - 0.02 µg/mL is recommended. It is recommended that the reagent be titrated for optimal performance for each application.
- RRID
-
AB_2728601 (BioLegend Cat. No. 852901)
AB_2728602 (BioLegend Cat. No. 852902)
Antigen Details
- Structure
- Human BACE1 is a 501 amino acid protein with a molecular mass of 56 kD and exists as 4 isoforms.
- Distribution
-
Tissue sources: Ubiquitously expressed. Highest levels in the brain and pancreas.
Cellular distribution: Plasma membrane, endoplasmic reticulum, and golgi apparatus. - Function
- BACE1 is involved in proteolytic processing of the amyloid precursor protein (APP). It cleaves APP at the N-terminus of the amyloid β peptide sequence, between residues 671 and 672 of APP.
- Interaction
- APP, IL1-R2, PSGL-1, ST6Gal-1, GGA1, GGA2, GGA3, RTN3, RTN4, SNX6, PCSK9, NAT8 and NAT8B.
- Biology Area
- Cell Biology, Neurodegeneration, Neuroscience, Protein Misfolding and Aggregation
- Molecular Family
- Enzymes and Regulators, Secretases
- Antigen References
-
- Vassar R, et al. 2014. J. Neurochem. 130:4-28.
- Kandalepas PC, Vassar R. 2014. Curr. Alzheimers Res. 11:441-9.
- Kandalepas PC, Vassar R. 2012. J. Neurochem. 120(Suppl. 1):55-61.
- Gruninger-Leitch F, et al. 2002. J. Biol. Chem. 277:4687-93.
- Koelsch G. 2017. Molecules. 22(10): E1723.
- Gene ID
- 23621 View all products for this Gene ID
- UniProt
- View information about BACE1 on UniProt.org
Related Pages & Pathways
Pages
Related FAQs
Other Formats
View All BACE1 Reagents Request Custom Conjugation| Description | Clone | Applications |
|---|---|---|
| Purified anti-BACE1 | A17035K | WB,Direct ELISA |
Compare Data Across All Formats
This data display is provided for general comparisons between formats.
Your actual data may vary due to variations in samples, target cells, instruments and their settings, staining conditions, and other factors.
If you need assistance with selecting the best format contact our expert technical support team.
 Login/Register
Login/Register 








Follow Us