- Regulatory Status
- RUO
- Other Names
- 3-chymotrypsin-like protease, 3CLpro, 3CL pro, 3CL-pro, main protease, M pro, M-pro
- Ave. Rating
- Submit a Review
- Product Citations
- publications
-
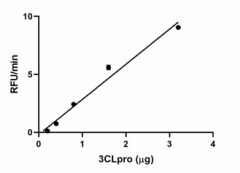
SARS-CoV-2 3C-like main protease (3CLpro) activity is measured by its ability to cleave the fluorogenic peptide substrate Dabcyl-KTSAVLQSGFRKME(Edans)-NH2. The accumulation of the cleavage product is quantified by the increase in fluorescence at 490 nm upon excitation at 340 nm. The SA is > 1.2 pmol/min/µg. -
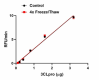

Stability Testing for Recombinant SARS-CoV-2 3C-Like Main Protease. Recombinant SARS-CoV-2 3C-like main protease was aliquoted in 50 mM HEPES, pH 7.5, 150 mM NaCl, 1mM DTT, 1 mM EDTA, 10% glycerol. One aliquot was frozen and thawed four times (4x Freeze/Thaw) and compared to the control that was kept at 4⁰C (Control). The samples were tested for their ability to cleave the fluorogenic peptide substrate Dabcyl-KTSAVLQSGFRKME(Edans)-NH2. The accumulation of the cleavage product was quantified by the increase in fluorescence at 490 nm upon excitation at 340 nm. The SA is > 1.2 pmol/min/µg.
| Cat # | Size | Price | Save |
|---|---|---|---|
| 795404 | 25 µg | ¥41,000 | |
| 795406 | 100 µg | ¥99,310 |
SARS CoV-2 3C-like main protease (3CLpro) is one of the two cysteine proteases that process the viral polyproteins of the SARS-CoV-2 virus responsible for the disease known as COVID-19 (1). Polyproteins pp1a and pp1ab are cleaved by 3CLpro and papain-like protease (PLpro) to produce 16 putative non-structural proteins. This proteolytic cleavage step is essential for viral replication and infection. 3CLpro is homodimeric in its active form and inactive as a monomer (2). The active form recognizes and cleaves the recognition sequence Leu-Gln↓(Ser/Ala/Gly). Domains I and II of 3CLpro consists of two β-barrel domains that form the chymotrypsin structure, and domain III contains a Cys-His catalytic dyad. The substrate binding site can be found in a cleft between domains I and II. 3CLpro, PLpro, and ACE2 currently serve as major therapeutic drug targets. There exists a variety of inhibitors of other coronavirus 3C-like proteases that may be considered for their antiviral potential against the novel coronavirus (3). These include small molecule inhibitors as well as larger molecules that mimic the recognition peptide structure. Inhibitors of the HIV Asp protease have been shown to treat SARS infection and are theorized to be effective inhibitors of 3CLpro (4). Some broad spectrum 3CLpro inhibitors have been shown to be effective against the SARS-CoV-2 in vitro (5).
Product DetailsProduct Details
- Source
- SARS CoV-2 3C-like main protease, amino acid (Ser3264 - Gln3569) (Accession: QHD43415.1) was expressed in E.coli. The N-terminal contains a GST-tag.
- Molecular Mass
- The 532 amino acid recombinant protein has a predicted molecular mass of approximately 60 kD. The protein migrates at approximately 60 kD by SDS-PAGE in both DTT-reduced and non-reduced conditions. The predicted N-terminal amino acid is Met.
- Purity
- > 96%, as determined by Coomassie stained SDS-PAGE
- Formulation
- 0.22 µm filtered protein solution is in 50mM HEPES, pH 7.5, 150mM NaCl, 1mM EDTA, 5mM TCEP, 10% Glycerol.
- Endotoxin Level
- Less than 0.1 EU per µg cytokine as determined by the LAL method
- Concentration
- 10 and 25 µg sizes are bottled at 200 µg/mL. 100 µg size and larger sizes are lot-specific and bottled at the concentration indicated on the vial. To obtain lot-specific concentration and expiration, please enter the lot number in our Certificate of Analysis online tool.
- Storage & Handling
- Unopened vial can be stored at -20°C or -70°C for six months. For maximum results, quick spin vial prior to opening. Avoid repeated freeze/thaw cycles.
- Activity
- SARS Cov-2 3C-like main protease activity is measured by the cleavage of fluorogenic peptide substrate Dabcyl-KTSAVLQSGFRKME(Edans)-NH2. Accumulation of the cleavage product is measured by fluorescence at 490 nm upon excitation at 340 nm. The specific activity is > 1.2 pmol/min/µg.
- Application
-
Bioassay
- Application Notes
-
SARS CoV-2 3C-Like Main Protease Assay Procedure
SARS CoV-2 3C-like main protease (3CLpro) activity is measured by its ability to cleave the fluorogenic peptide substrate Dabcyl-KTSAVLQSGFRKME(Edans)-NH2. The accumulation of the cleavage product is quantified by the increase in fluorescence at 490 nm upon excitation at 340 nm.
Materials and Buffers
- Assay buffer: 20 mM Tris-HCl, 150 mM NaCl, pH 8.0, 5 mM DTT
- 3CLpro substrate: Dabcyl-KTSAVLQSGFRKME(Edans)-NH2 (CPC Scientific Cat. No. COVD-001)
- Recombinant 3CLpro
- F16 Black Maxisorp plate (Nunc, Cat. No. 475515)
- Fluorescent plate reader (Model: SpectraMax Gemini EM by Molecular Devices) or equivalent
Assay Procedure
- Dilute the substrate to 20 µM in the assay buffer.
- Dilute the recombinant 3CLpro to 64 µg/mL. Make six 1:2 dilutions in a black well plate. Include a blank with 50 µL assay buffer.
- Start the reaction by adding 50 µL of the diluted substrate to each well.
- Take a baseline reading at excitation and emission wavelengths of 340 nm and 490 nm (top read), respectively, in endpoint mode.
- Cover plate and incubate in 37°C. Take additional readings at 30 minutes and 60 minutes after the substrate addition.
- Calculate the specific activity using the slope of total RFU per µg enzyme, obtained from either the 30 minute or 60 minute reading:
Specific Activity (pmol/min/µg) = Slope* (RFU/µg enzyme) x Conversion Factor** (pmol/RFU)
Time (min)
*Adjusted for substrate blank **Derived using calibration standard EDANS (Ana Spec, Cat. No. AS-23886)Per well:- 3CLpro: 3.2, 1.6, 0.8, 0.4, 0.2, 0.1, 0.05 µg
- Substrate: 10 µM
BioLegend carrier-free recombinant proteins provided in liquid format are shipped on blue-ice. Our comparison testing data indicates that when handled and stored as recommended, the liquid format has equal or better stability and shelf-life compared to commercially available lyophilized proteins after reconstitution. Our liquid proteins are validated in-house to maintain activity after shipping on blue ice and are backed by our 100% satisfaction guarantee. If you have any concerns, contact us at tech@biolegend.com.
Antigen Details
- Structure
- Dimer
- Distribution
-
SARS-CoV-2 virus, 3CLpro is the main protease found in coronaviruses
- Function
- Proteolytic processing of viral polyproteins pp1a and pp1ab
- Interaction
- Peptides containing L-Q-(S/A/G)
- Bioactivity
- Cleaves polypeptide substrate Dabcyl-KTSAVLQSGFRKME(Edans)-NH2
- Antigen References
-
1. Dömling A & Gao L, 2020. Chem. 6:1283.2. Pillaiyar T, et al. 2016. J. Med. Chem. 59:6595.3. De Clercq E, 2006. Expert Rev. Anti. Infect. Ther. 4:291.4. Chu CM, et al. 2004. Thorax. 59:252.5. Rathnayake AD, et al. 2020. Sci. Transl. Med. 12:eabc5332.6. Cui J, et al. 2019. Nat. Rev. Microbiology. 17:181.
7. Douangamath A, et al. 2020. Nat. Comun. 11:5047 - Gene ID
- 43740578 View all products for this Gene ID
- UniProt
- View information about 3C-Like Main Protease on UniProt.org
Related Pages & Pathways
Pages
Related FAQs
- Why choose BioLegend recombinant proteins?
-
• Each lot of product is quality-tested for bioactivity as indicated on the data sheet.
• Greater than 95% Purity or higher, tested on every lot of product.
• 100% Satisfaction Guarantee for quality performance, stability, and consistency.
• Ready-to-use liquid format saves time and reduces challenges associated with reconstitution.
• Bulk and customization available. Contact us.
• Learn more about our Recombinant Proteins. - How does the activity of your recombinant proteins compare to competitors?
-
We quality control each and every lot of recombinant protein. Not only do we check its bioactivity, but we also compare it against other commercially available recombinant proteins. We make sure each recombinant protein’s activity is at least as good as or better than the competition’s. In order to provide you with the best possible product, we ensure that our testing process is rigorous and thorough. If you’re curious and eager to make the switch to BioLegend recombinants, contact your sales representative today!
- What is the specific activity or ED50 of my recombinant protein?
-
The specific activity range of the protein is indicated on the product datasheets. Because the exact activity values on a per unit basis can largely fluctuate depending on a number of factors, including the nature of the assay, cell density, age of cells/passage number, culture media used, and end user technique, the specific activity is best defined as a range and we guarantee the specific activity of all our lots will be within the range indicated on the datasheet. Please note this only applies to recombinants labeled for use in bioassays. ELISA standard recombinant proteins are not recommended for bioassay usage as they are not tested for these applications.
- Have your recombinants been tested for stability?
-
Our testing shows that the recombinant proteins are able to withstand room temperature for a week without losing activity. In addition the recombinant proteins were also found to withstand four cycles of freeze and thaw without losing activity.
- Does specific activity of a recombinant protein vary between lots?
-
Specific activity will vary for each lot and for the type of experiment that is done to validate it, but all passed lots will have activity within the established ED50 range for the product and we guarantee that our products will have lot-to-lot consistency. Please conduct an experiment-specific validation to find the optimal ED50 for your system.
- How do you convert activity as an ED50 in ng/ml to a specific activity in Units/mg?
-
Use formula Specific activity (Units/mg) = 10^6/ ED50 (ng/mL)








Follow Us