Governing the Fate of Stem Cells With Transcription Factors
Stem cells are unique due to their ability to limitlessly self-renew and differentiate into each cell type in the adult body. The capability of these cells to differentiate depends on the stem cell type, the regulation of gene expression by various transcription factors and interaction with the stem cell niche1,4.
Our scientists have developed a wide array of stem cell-focused reagents and resources for many applications including flow cytometry, western blotting, ELISAs, and recombinant proteins for cell differentiation.
Transcription factors are proteins that regulate the transcription of genes, or the production of mRNA from DNA. Their unique feature is their ability to bind DNA at sequences known as promoter or enhancer regions. Once the transcription factor binds to an enhancer region, this can cause stimulation or repression of gene transcription. Transcription factors have an important role in the ability of a cell to self-renew and also differentiate into most cell types, also known as pluripotency1. View our full list of available antibodies for transcription factors.
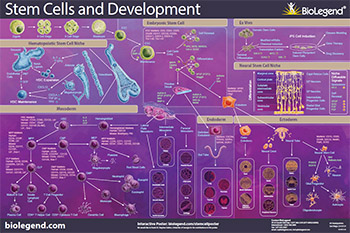
Download or request our Stem Cells and Development poster to explore the pathways of stem cell biology including differentiation from the mesoderm, endoderm, and ectoderm germ layers, iPS cells, and interactions with their niche.
Rb
Rb, or retinoblastoma protein, is a key regulator of the cell cycle, particularly during the transition from the G1 to S phases. The underphosphorylated, active form of Rb interacts directly with E2F1, leading to cell cycle arrest, while the hyperphosphorylated form decouples from E2F1, thus promoting the transcription of genes promoting entry into the S phase. Rb is directly involved in heterochromatin formation by maintaining chromatin structure and that of constitutive heterochromatin by stabilizing histone methylation. Rb homeostasis is also essential for self-renewal and survival of human embryonic stem cells10.
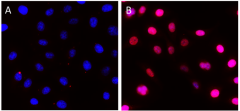
HeLa cells were fixed, permeabilized, blocked and intracellularly stained with either Purified Rat IgG2a, κ Isotype Ctrl Antibody (Panel A) or Purified anti-Rb1 Antibody (Panel B) and Alexa Fluor® 594 secondary antibody. Nuclei were counterstained with DAPI.
RUNX1
RUNX1 belongs to the runt domain family of transcription factors and regulates target gene expression through forming a heterodimeric DNA-binding complex with CBFB. RUNX1 was first identified as a RUNX1-ETO fusion protein in acute myeloid leukemia (AML) and is frequently mutated in AML and myelodysplastic syndrome due to chromosomal translocation. RUNX1-deficient mice fail to generate hematopoietic stem cells. RUNX1 regulates CD4 gene transcription during multiple stages of T cell development and represses the CD4 gene in CD4-CD8- (double negative) T cells. RUNX1 is also required for the differentiation of CD8+, Th17, and regulatory T cells. Impaired differentiation of megakaryocytes and decreased circulating platelet count were observed in the absence of RUNX1. These findings revealed that RUNX1 acts as a tumor suppressor for myeloid leukemia and is crucial for the development and terminal differentiation of several blood cell lineages2,3.
Jurkat cells were stained with purified anti-RUNX1 antibody, followed by staining with DyLight™ 488 conjugated secondary IgG antibody (green). Nuclei stained with DAPI (blue).
Pax6
Pax6 is a transcription factor present during embryonic development. The encoded protein contains two different binding sites that are known to bind DNA and function as a regulator of gene transcription. It is a key regulatory gene of eye and brain development. Within the brain, the protein is involved in the development of specialized cells that process smell. Pax-6 acts as a critical gene for the development of eyes and other sensory organs, certain neural and epidermal tissues as well as other homologous structures, usually derived from ectodermal tissues. Pax6 serves as a regulator in the coordination and pattern formation required for differentiation and proliferation to successfully take place, ensuring that the processes of neurogenesis and oculogenesis are carried out successfully. As a transcription factor, Pax6 acts at the molecular level in the signaling and formation of the central nervous system11.
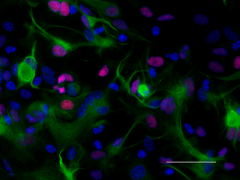
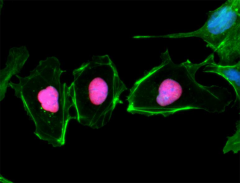
ICC staining of purified anti-Pax6 antibody (clone AD2.35) on mouse primary astrocytes. The cells were fixed, permeabilized, blocked and then stained with primary antibody, and Alexa Fluor® 594 conjugated secondary IgG (Cat. No. 405326). Alexa Fluor® 488 conjugated anti-GFAP antibody (clone SMI 25) was used to stain astrocytes. Nuclei were counterstained with DAPI.
STAT3
STAT3 protein belongs to a group of intracellular transcription factors that mediate a variety of functions such as cellular differentiation, proliferation, and apoptosis. STAT3 is activated by a number of extracellular signaling proteins such as growth factors VEGF and EGF. The binding of these factors to their respective receptors VGFR and EGFR initiates signaling cascades that involve phosphorylation and activation of protein kinases in the cytoplasm, which in turn can phosphorylate STAT3. It forms homodimers with itself or heterodimers with STAT1 when tyrosine is phosphorylated, and then translocates to the nucleus, acting as a transcription regulator. Once in the nucleus, STAT3 can bind to interferon-stimulated response element (IRSE) or gamma interferon-activated sequence (GAS) promoter regions, and together with co-factors such as Nuclear Factor Kappa (NF-κB) and hypoxia-inducible factor 1 alpha (HIF-1α), can induce transcription of genes that are involved in inflammation, cell proliferation, and survival8.
Due to the tumorigenic potential of STAT3, these factors could trigger aberrant cell growth and cell cycle transition, angiogenesis, invasion, and tumor metastasis, which leads to modulation of the expression of the proteins involved in these pathways8. Some examples are alterations in the expression of the following proteins: B-cell lymphoma 2 (Bcl-2) an anti-apoptotic protein that regulates mitochondrial dynamics, cyclins D1-3 and cyclin A, proteins involved in G1 to S phase transition, and matrix metalloproteinase 9 (MMP-9) a protein that breaks down the extracellular matrix, contributing to invasion and metastasis.
Transcription Factors and Pluripotency
Induced pluripotent stem cells, or iPS cells, are adult somatic cells that have been “de-differentiated” and resemble embryonic stem cells. First described in 2006, Yamanaka and Takahashi reported that expression of only four factors, Oct-4, SOX2, Klf4, and MYC, could reprogram somatic cells into pluripotent stem cells that are able to differentiate into all three germ layers4. Co-expression of SOX with OCT4, MYC, and KLF4 is sufficient to reprogram somatic cells to induced pluripotent stem cells, which exert similar characteristics as natural pluripotent stem cells.
SOX2
SOX2 is the most studied member of the SRY-related box transcription factor family. It binds to target genes through its highly conserved HMG box domain. Inactivation of the SOX2 gene causes lethality during embryonic development. SOX2 knockdown in embryonic stem cells results in their differentiation. These findings indicate that SOX2 is crucial for the self-renewal and pluripotency of embryonic stem cells. In addition, over-expression of SOX2 has been found in various types of malignant cancer. Knockdown of SOX2 results in cell cycle arrest by downregulating cyclin D1 and inhibition of tumor cell proliferation, suggesting that SOX2 is involved in activating genes associated with tumor progression2.
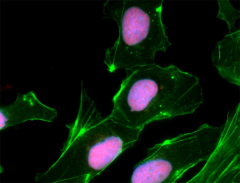
NTERA-2 cells were fixed, permeabilized, blocked and intracellularly stained with SOX2 Antibody (clone 14A6A34) followed by DyLight™ 594 secondary IgG (red). Actin filaments were labeled with Alexa Fluor® 488 Phalloidin (green). Nuclei were counterstained with DAPI (blue).
Oct4
Oct4 (Octamer binding transcription factor 4) is an important marker of the undifferentiated state and central regulator of pluripotency in ES cells. When embryonic stem cells are triggered to differentiate, Oct4 is downregulated, thus providing a model for the early events linked to somatic differentiation in the developing embryo.
NTERA-2 cells were fixed, permeabilized, blocked and intracellularly stained with anti-Oct4 (Oct3) (3A2A20) and followed by DyLight™ 594 secondary IgG (red). Actin filaments were labeled with Alexa Fluor® 488 Phalloidin (green). Nuclei were counterstained with DAPI (blue).
There have been many advancements in reprogramming techniques including modifying both the factors used and how they are delivered to the cells. For example, some studies have found that other combinations of transcription factors may be able to induce reprogramming (including Nanog and Lin-28). Because viral transduction methods require incorporating these genes into the genome, other methods of reprogramming have also been studied including using small molecule compounds, introducing transcription factors through modified mRNAs, and CRISPR/Cas9 editing9.
Applications
Stem cells can be used to create disease models to learn about the molecular mechanisms of disease and to test novel drug molecules. Stem cells can also be used in the lab to generate many other types of cells, including dopamine cells5.
In late 2017, researchers in Jun Takahashi’s lab at the Center for iPS Cell Research and Application at Kyoto University announced that a study transplanting nerve cells made from iPS cells into the brains of pre-clinical models was promising. The grafted cells were able to secrete dopamine and stimulate neurons in the brain and survived for two years, appeared to improve symptoms, and did not cause ill side effects6.
From Stoker, Thomas B. "Stem cell treatments for Parkinson’s disease." Exon Publications (2018): 161-175. Licensed under Creative Commons Attribution 4.0 International (CC BY 4.0).
A clinical trial to move the procedure into humans began in 20187. Dopaminergic progenitor cells which develop into neurons that produce dopamine were injected directly into an area of the brain associated with neural degeneration in Parkinson’s disease. A positive response was observed but further studies are required to understand long-term effects.
The capacity for stem cells to differentiate into any cell type has created a large interest in various applications and is an active area of investigation. Understanding the complexities of stem cell differentiation and development can reveal novel ways to study wound and tissue repair, oncology, and neurodegenerative diseases. As the field advances, our scientists are committed to developing resources and reagents for your stem cell research.
References
- Patterson, Michaela et al. "Defining the Nature of Human Pluripotent Stem Cell Progeny." Cell research vol. 22,1 (2012): 178–93 (2012). doi: 10.1038/cr.2011.133
- Friebel, Ekaterina et al. "Single-Cell Mapping of Human Brain Cancer Reveals Tumor-Specific Instruction of Tissue-Invading Leukocytes." Cell vol. 181,7 (2020): 1626-1642.e20. doi:10.1016/j.cell.2020.04.055
- Goyama, Susumu et al. "Transcription factor RUNX1 promotes survival of acute myeloid leukemia cells." The Journal of clinical investigation vol. 123,9 (2013): 3876-88. doi: 10.1172/JCI68557
- Takahashi, Kazutoshi et al. "Induction of pluripotent stem cells from adult human fibroblasts by defined factors." Cell vol. 131,5 (2007): 861-72. doi: 10.1016/j.cell.2007.11.019
- Barker, Roger A et al. Cell-based therapies for Parkinson disease—past insights and future potential. Nature reviews. Neurology vol. 11,9 (2015): 492–503. doi: 10.1038/nrneurol.2015.123
- Stoker, Thomas Benjamin. "Stem Cell Treatments for Parkinson’s Disease." Parkinson's Disease: Pathogenesis and Clinical Aspects, edited by Thomas B. Stoker et. al., Codon Publications, 21 December 2018. doi:10.15586/codonpublications.parkinsonsdisease.2018.ch9
- Cyranoski, David. "'Reprogrammed’ stem cells implanted into patient with Parkinson’s disease." Nature vol. 563 (2018): 1-2. doi: 10.1038/d41586-018-07407-9
- Loh, Chin-Yap et al. "Signal Transducer and Activator of Transcription (STATs) Proteins in Cancer and Inflammation: Functions and Therapeutic Implication." Frontiers in oncology vol. 9 48. Feb. 2019, doi:10.3389/fonc.2019.00048
- Takahashi, Kazutoshi, and Shinya Yamanaka. "A decade of transcription factor-mediated reprogramming to pluripotency." Nature Reviews. Molecular cell biology vol. 17,3 (2016): 183–93. doi: 10.1038/nrm.2016.8
- Conklin, Jamie F et al. The RB family is required for the self-renewal and survival of human embryonic stem cells." Nature communications vol. 3 (2012): 1244. doi: 10.1038/ncomms2254
- Chi, Neil, and Jonathan A Epstein. "Getting your Pax straight: Pax proteins in development and disease." Trends in genetics : TIG vol. 18,1 (2002): 41-7. doi: 10.1016/S0168-9525(01)02594-X







Follow Us