- Clone
- 3F4 (See other available formats)
- Regulatory Status
- RUO
- Other Names
- Major prion protein, p27-30, CD230 antigen, prion protein PrP, prion-related protein
- Previously
-
Signet Catalog# 9640-02
Signet Catalog# 9640-05
Signet Catalog# 9640-10
Covance Catalog# SIG-39640
- Isotype
- Mouse IgG2a, κ
- Ave. Rating
- Submit a Review
- Product Citations
- publications
-

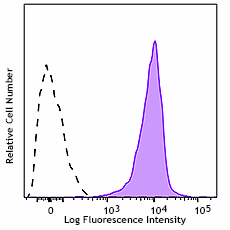
Human peripheral blood lymphocytes were stained with Biotin anti-CD230 (Prion) antibody (clone 3F4, filled histogram) or Alexa Fluor™ 647 Mouse IgG2a, κ isotype control antibody (Cat. No. 400234) (open histogram). The cells that were incubated with Biotin anti-CD230 (Prion) antibody only were stained with Alexa Fluor™ 647 Streptavidin (Cat. No. 405237).
Prions cause neurodegenerative disease by aggregating extracellularly within the central nervous system which disrupt the normal tissue structure. This disruption is characterized by "holes" in the tissue with resultant spongy architecture. Two conformational isoforms exist, the normal cellular isoform (PrPC) and the infectious, scrapie isoform (PrPSC). Other histological changes include astrogliosis and the absence of an inflammatory reaction. Neurodegenerative symptoms can include convulsions, dementia, ataxia (balance and coordination dysfunction), and behavioral or personality changes.
All known prion diseases are collectively called transmissible spongiform encephalopathies (TSEs). Prion (PrP) is highly conserved through mammals and comparison between primates ranges from 92.9-99.6% similarity in amino acid sequence. The human protein structure consists of a globular domain with three α-helices and a two-strand antiparallel β-sheet, an NH2-terminal tail, and a short COOH-terminal tail. A glycosylphosphatidylinositol (GPI) membrane anchor at the COOH-terminal tethers PrP to cell membranes. This anchor is integral to the transmission of conformational change; secreted PrP lacking the anchor component is unaffected by the infectious isoform. PrPSC accumulates in compact, protease-resistant aggregates within neural tissue and has a different secondary and tertiary structure from PrPC, but an identical primary sequence.
The primary sequence of PrP is 253 amino acids long before posttranslational modification. Signal sequences in the amino- and carboxy- terminal ends are removed posttranslationally, resulting in a mature length of 208. For human and Syrian hamster PrP, two glycosylated sites exist on helices 2 and 3 at Asn181 and Asn197. Murine PrP has glycosylation sites as Asn180 and Asn196. A disulfide bond exists between Cys179 of the second helix and Cys214 of the third helix (human PrPC numbering).
The precise function of PrP is not yet known, but it is possibly involved in the transport of ionic copper to cells from the surrounding environment. Researchers have also proposed roles for PrP in cell signaling or in the formation of synapses.
Spatial learning, a predominantly hippocampal-function, is decreased in PrP null mice and can be recovered with the reinstatement of PrP in neurons; indicating that loss of PrP function is the cause. PrP is present in both pre- and post-synaptic neuron cells, and the greatest concentration is in the pre-synaptic cells. Some research indicates PrP involvement in neuronal development, differentiation, and neurite outgrowth. The PrP-activated signal transduction pathway is associated with axon and dendritic outgrowth with a series of kinases.
Though most attention is focused on PrP’s presence in the nervous system, it is also abundant in immune system tissue. PrP immune cells include haematopoietic stem cells, mature lymphoid and myeloid compartments, and certain lymphocytes; also, it has been detected in natural killer cells, platelets, and monocytes. T cell activation is accompanied by a strong up-regulation of PrP, though it is not requisite. The lack of immuno-response to transmissible spongiform encephalopathies (TSE), neurodegenerative diseases caused by prions, could stem from the tolerance for PrPSc.
Product Details
- Verified Reactivity
- Human
- Antibody Type
- Monoclonal
- Host Species
- Mouse
- Formulation
- Phosphate-buffered solution.
- Preparation
- The antibody was purified by affinity chromatography.
- Concentration
- 1.0 mg/ml
- Storage & Handling
- The antibody solution should be stored undiluted between 2°C and 8°C. Please note the storage condition for this antibody has been changed from -20°C to between 2°C and 8°C. You can also check your vial or your CoA to find the most accurate storage condition for this antibody.
- Application
-
FC - Quality tested
- Recommended Usage
-
Each lot of this antibody is quality control tested by immunofluorescent staining with flow cytometric analysis. For flow cytometric staining, the suggested use of this reagent is ≤ 1.25 - 5.0 µg per million cells in 100 µl volume. It is recommended that the reagent be titrated for optimal performance for each application.
- Application Notes
-
3F4 is reactive in amino acid residues 109-112 of prion protein (PrP) from humans, hamsters, and felines. It recognizes both protease sensitive and protease resistant forms of PrP (after denatured).
This antibody is exclusively provided by BioLegend. -
Application References
(PubMed link indicates BioLegend citation) -
- Kondoh G, et al. 2005. Nat. Med. 11(2):160-6. (IHC-F)
- Gilch S, et al. 2004. Traffic. 5(4):300-13. (WB, IHC-F, ELISA) PubMed
- Nishina K, et al. 2004. Biochemistry 43:2613-21. (IP)
- Shiga Y, et al. 2004. Neurology 63(3):443-9. (WB, IHC-F, IP, ELISA)
- Zou WQ, et al. 2004. Proc. Natl. Acad. Sci. USA 101(5):1380-5.
- Kitamoto T, et al. 1987. Lab Invest. 57(2):230-6.
- Kascsak RJ, et al. 1987. J. Virol. 61(12):3688-93.
- Product Citations
-
- RRID
-
AB_2564625 (BioLegend Cat. No. 800306)
AB_2564627 (BioLegend Cat. No. 800305)
AB_2564626 (BioLegend Cat. No. 800304)
Antigen Details
- Biology Area
- Cell Biology, Neurodegeneration, Neuroscience, Protein Misfolding and Aggregation
- Molecular Family
- Prion (CD230)
- Antigen References
-
1. Asante EA, et al. 2015. Nature 522:478.
2. Mead S, et al. 2013. N. Engl. J. Med. 369:1904.
3. Chakrabarti O, Hegde RS. 2009. Cell 137:1136.
4. Wadsworth JD, et al. 2004. Science 306:1793. - Gene ID
- 5621 View all products for this Gene ID
- UniProt
- View information about CD230 on UniProt.org
Related Pages & Pathways
Pages
Related FAQs
- How many biotin molecules are per antibody structure?
- We don't routinely measure the number of biotins with our antibody products but the number of biotin molecules range from 3-6 molecules per antibody.
Other Formats
View All CD230 (Prion) Reagents Request Custom Conjugation| Description | Clone | Applications |
|---|---|---|
| Purified anti-CD230 (Prion) | 3F4 | FC,ELISA,WB,IHC-F,IP |
| Anti-CD230 (Prion) | 3F4 | FC |
| Biotin anti-CD230 (Prion) | 3F4 | FC |
| FITC anti-human CD230 (Prion) | 3F4 | FC |
| APC anti-human CD230 (Prion) | 3F4 | FC |
| PE anti-human CD230 (Prion) | 3F4 | FC |
Compare Data Across All Formats
This data display is provided for general comparisons between formats.
Your actual data may vary due to variations in samples, target cells, instruments and their settings, staining conditions, and other factors.
If you need assistance with selecting the best format contact our expert technical support team.
-
Purified anti-CD230 (Prion)
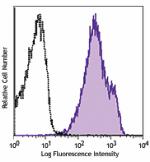
Human peripheral blood lymphocytes were stained with purifie... -
Anti-CD230 (Prion)
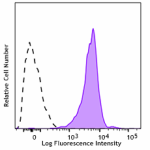
Human peripheral blood lymphocytes were stained with anti-CD... -
Biotin anti-CD230 (Prion)
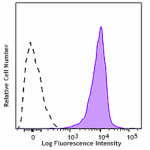
Human peripheral blood lymphocytes were stained with Biotin ... -
FITC anti-human CD230 (Prion)
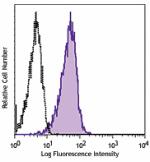
Human peripheral blood lymphocytes were stained with CD230 (... -
APC anti-human CD230 (Prion)
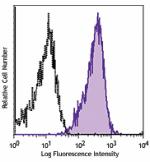
Human peripheral blood lymphocytes were stained with CD230 (... -
PE anti-human CD230 (Prion)
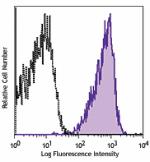
Human peripheral blood lymphocytes were stained with CD230 (...
 Login / Register
Login / Register 












Follow Us