- Clone
- 16B12 (See other available formats)
- Regulatory Status
- RUO
- Other Names
- Hemagglutinin tag, HA tag
- Previously
-
Covance Catalog# AFC-101P
- Isotype
- Mouse IgG1, κ
- Ave. Rating
- Submit a Review
- Product Citations
- publications
| Cat # | Size | Price | Quantity Check Availability | Save | ||
|---|---|---|---|---|---|---|
| 900801 | 1 mL | $633 | ||||
The HA tag (hemagglutinin) is an amino acid sequence derived from the human influenza hemagglutinin surface glycoprotein, corresponding to amino acids 98-106. It is commonly used as a tag to facilitate detection, isolation, and purification of proteins. The full amino acid sequence is: YPYDVPDYA.
Product DetailsProduct Details
- Verified Reactivity
- Influenza, Epitope tag
- Reported Reactivity
- Species independent
- Antibody Type
- Monoclonal
- Host Species
- Mouse
- Immunogen
- Monoclonal antibody HA.11 was raised against the twelve amino acid peptide CYPYDVPDYASL.
- Formulation
- Purified IgG immobilized on Sepharose™ Fast Flow beads (in PBS + 0.03% Thimerosal).
- Preparation
- The antibody was purified by affinity chromatography.
- Storage & Handling
- Store between 2-8°C. The matrix may be re-used several times. To strip column after use, wash with several bead volumes of 0.1 M glycine pH 2.8 followed immediately by PBS containing 0.3% Thimerosal or 1mM Sodium Azide as preservative.
- Application
-
Purification, IP
- Recommended Usage
-
Each lot of this antibody is quality control tested by immunoprecipitation.
The optimal buffers and matrix concentration should be determined for each specific assay condition.
Binding: Tagged protein will bind to matrix in common physiologic buffers with pH in the range of 6.0-7.5, salt from 50-500 mM and in the presence of reasonable levels of detergent. Excess reducing agent should be avoided as the disulfide bridges holding antibody heavy and light chains may be compromised. BioLegend tests Affinity Matrix using an equilibration/binding buffer containing:
• 100 mM Tris-HCl (pH 7.5)
• 150 mM NaCl
• 0.1% Tween 20
• 0.5% BSA
• 1 mM beta-mercaptoethanol
Washing: After binding, washes with several bead volumes of buffer are recommended. Such buffer may contain increased salt, altered pH, etc. as determined empirically to remove un-tagged proteins.
Elution: Several options are available for elution.
1. SDS gel loading buffer may be applied directly to the beads in order to display all bound protein on a polyacrylamide gel/western. Note that gel loading buffer containing reducing agent will also release some antibody heavy and light chains (approx 25 and 50kD, respectively).
2. Competitive elution with epitope peptide. For epitope tag affinity matrices, prepare an elution buffer with epitope tag peptide at 400 ug/mL in 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, 1 mM EDTA (pH 8.0).
3. Chemical Elution. Elution by pH or chaotropic salts is also possible. For elution by pH, either 0.1 M glycine pH 2.8 or 40 mM diethyl-amine pH 11.0 may be used. - Application Notes
-
This affinity matrix can be used for immunopurification of HA-tagged fusion proteins from crude starting material. On an analytical scale, it can also be used for immunoprecipitating HA-tagged proteins.
Monoclonal mouse antibody HA.11 recognizes the peptide epitope, YPYDVPDYA. This second-generation HA antibody is an excellent substitute for the 12CA5 monoclonal antibody. The HA.11 antibody recognizes HA epitopes located in the middle of protein sequences as well as at the N- or C-terminus.
HA.11 antibody was purified using protein-G chromatography and was subsequently immobilized onto a Sepharose™ Fast Flow matrix.
Sepharose is a trademark of Amersham Biosciences Limited -
Application References
(PubMed link indicates BioLegend citation) -
- Ferrando A, et al. 2001. Nucleic Acids Res. 29:3685.
- J Field, et al. 1988. Mol Cell Biol. 8:2159.
- Bennett BD, et al. 2000. J Biol Chem. 275:37712. (IF, IP, WB) PubMed
- Product Citations
-
- RRID
-
AB_2564999 (BioLegend Cat. No. 900801)
Antigen Details
- Biology Area
- Cell Biology
- Gene ID
- NA
- UniProt
- View information about HA.11 on UniProt.org
Related Pages & Pathways
Pages
Related FAQs
Other Formats
View All HA.11 Reagents Request Custom ConjugationCustomers Also Purchased

Compare Data Across All Formats
This data display is provided for general comparisons between formats.
Your actual data may vary due to variations in samples, target cells, instruments and their settings, staining conditions, and other factors.
If you need assistance with selecting the best format contact our expert technical support team.
-
Anti-HA.11 Epitope Tag Affinity Matrix
-
Alexa Fluor® 488 anti-HA.11 Epitope Tag
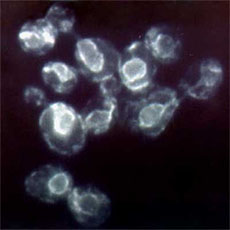
HA tagged CHO cells stained with Alexa Fluor® 488 Labeled An... 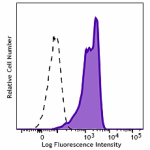
Transfected RBL1 cells expressing the HA tag on the cell sur... -
Alexa Fluor® 594 anti-HA.11 Epitope Tag
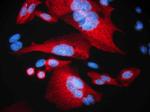
Staining of HA.11 Clone 16B12 Monoclonal Antibody, Alexa Flu... -
Anti-HA.11 Epitope Tag
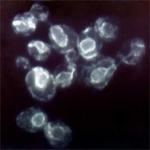
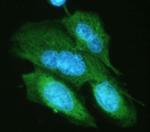
Immunofluorescence of HA.11 tagged Sbhlp protein. Photo cour... 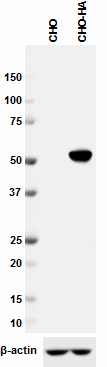
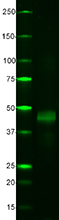
Western blot analysis of cell lysates from CHO and CHO HA st... 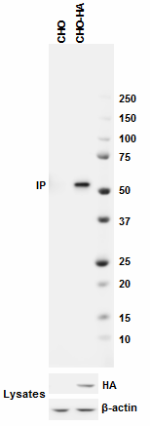
Immunoprecipitation of HA protein from CHO and CHO-HA stable...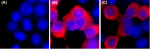 
Immunofluorescence staining of HeLa cells transfected withou... -
Biotin anti-HA.11 Epitope Tag
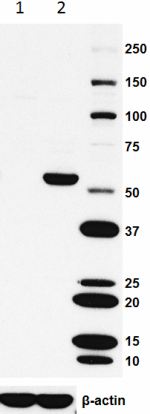
Total lysates (15 µg protein) from CHO (lane 1) and CHO HA s... 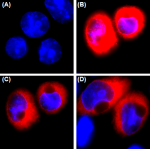
Immunofluorescence of HA-tagged plasmid-transfected A549 cel... -
FITC anti-HA.11 Epitope Tag
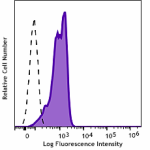
Transfected RBL1 cells expressing the HA tag on the cell sur... -
Purified anti-HA.11 Epitope Tag
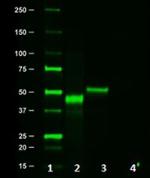
Western blot of Clone 16B12. Lane 1: Molecular weight marker... 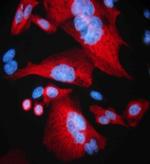
Staining of Clone 16B12 on methanol fixed CHO cells transfec... 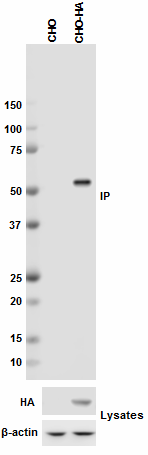
Immunoprecipitation of HA protein from CHO and CHO-HA stable... 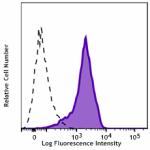
Transfected RBL1 cells expressing the HA tag on the cell sur... -
Alexa Fluor® 647 anti-HA.11 Epitope Tag
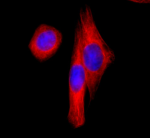
HA tag stably transfected CHO cells were fixed with ice cold... 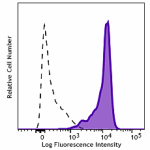
Transfected RBL1 cells expressing the HA tag on the cell sur... -
PE anti-HA.11 Epitope Tag
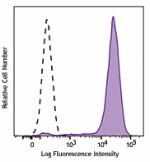
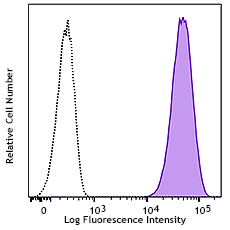
CHO-K1 cells (open histogram) or HA tag stably transfected c... 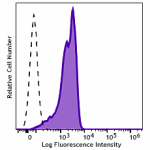
Transfected RBL1 cells expressing the HA tag on the cell sur... -
Direct-Blot™ HRP anti-HA.11 Epitope Tag
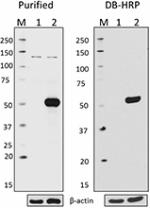
Total cell lysate from CHO (lane 1) and CHO stably transfect... -
Ultra-LEAF™ Purified anti-HA.11 Epitope Tag
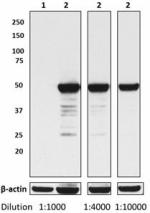
Total cell lysate (15 µg protein) from CHO (lane 1) and CHO ... 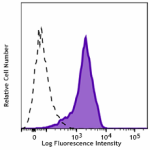
Transfected RBL1 cells expressing the HA tag on the cell sur... -
Brilliant Violet 421™ anti-HA.11 Epitope Tag
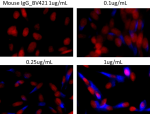
CHO cells stably transfected with HA tag were fixed with 4% ... 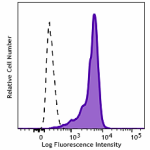
Transfected RBL1 cells expressing the HA tag on the cell sur... -
PE/Dazzle™ 594 anti-HA.11 Epitope Tag
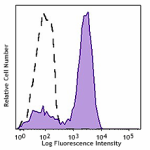
CHO-K1 cells (open histogram) or HA tag stably transfected c... 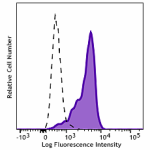
Transfected RBL1 cells expressing the HA tag on the cell sur... -
PE/Cyanine7 anti-HA.11 Epitope Tag
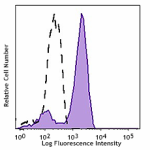
CHO-K1 cells (open histogram) or HA tag stably transfected c... 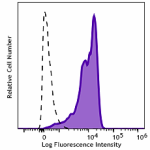
Transfected RBL1 cells expressing the HA tag on the cell sur... -
Pacific Blue™ anti-HA.11 Epitope Tag
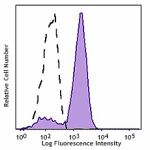
CHO-K1 cells (open histogram) or HA tag stably transfected c... 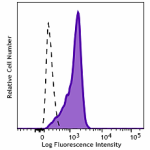
Transfected RBL1 cells expressing the HA tag on the cell sur... -
APC anti-HA.11 Epitope Tag
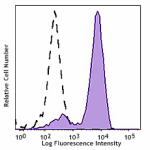
CHO-K1 cells (open histogram) or HA tag stably transfected c... 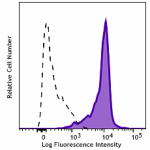
Transfected RBL1 cells expressing the HA tag on the cell sur... -
PerCP/Cyanine5.5 anti-HA.11 Epitope Tag
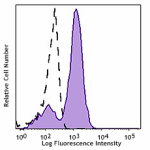
CHO-K1 cells (open histogram) or HA tag stably transfected c... 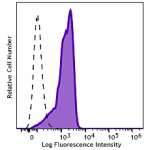
Transfected RBL1 cells expressing the HA tag on the cell sur... -
TotalSeq™-C1131 anti-HA.11 Epitope Tag
-
TotalSeq™-A1131 anti-HA.11 Epitope Tag
-
TotalSeq™-B1131 anti-HA.11 Epitope Tag
 Login/Register
Login/Register 







Follow Us