- Clone
- MF-14 (See other available formats)
- Regulatory Status
- RUO
- Other Names
- Forkhead box protein P3, Scurfin, JM2, IPEX, Zinc finger protein JM2
- Isotype
- Rat IgG2b, κ
- Ave. Rating
- Submit a Review
- Product Citations
- publications
-
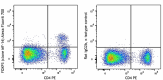

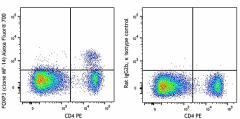
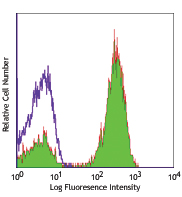
C57BL/6 splenocytes were surface stained with CD4 PE and then treated with True-Nuclear™ Transcription Factor Buffer Set. Cells were then stained with FOXP3 (clone MF-14) Alexa Fluor® 700 (left) or Rat IgG2b, κ Alexa Fluor® 700 isotype control (right).
| Cat # | Size | Price | Quantity Check Availability | Save | ||
|---|---|---|---|---|---|---|
| 126421 | 25 µg | $171 | ||||
| 126422 | 100 µg | $375 | ||||
FOXP3 is a 47 kD transcription factor, also known as Forkhead box protein P3, Scurfin, JM2, or IPEX. It is proposed to be a master regulatory gene and more specific marker of T regulatory cells than most cell surface markers (such as CD4 and CD25). Transduced expression of FOXP3 in CD4+/CD25- cells has been shown to induce GITR, CD103, and CTLA4 and impart a T regulatory cell phenotype. FOXP3 is mutated in X-linked autoimmunity-allergic dysregulation syndrome (XLAAD or IPEX) in humans and in "scurfy" mice. Overexpression of FOXP3 has been shown to lead to a hypoactive immune state suggesting that this transcriptional factor is a central regulator of T cell activity.
Product DetailsProduct Details
- Verified Reactivity
- Mouse
- Antibody Type
- Monoclonal
- Host Species
- Rat
- Formulation
- Phosphate-buffered solution, pH 7.2, containing 0.09% sodium azide.
- Preparation
- The antibody was purified by affinity chromatography and conjugated with Alexa Fluor® 700 under optimal conditions.
- Concentration
- 0.5 mg/ml
- Storage & Handling
- The antibody solution should be stored undiluted between 2°C and 8°C, and protected from prolonged exposure to light. Do not freeze.
- Application
-
ICFC - Quality tested
- Recommended Usage
-
Each lot of this antibody is quality control tested by intracellular flow cytometry using our True-Nuclear™ Transcription Factor Staining Protocol. For flow cytometric staining, the suggested use of this reagent is ≤0.06 µg per million cells in 100 µl volume. It is recommended that the reagent be titrated for optimal performance for each application.
* Alexa Fluor® 700 has a maximum emission of 719 nm when it is excited at 633 nm / 635 nm. Prior to using Alexa Fluor® 700 conjugate for flow cytometric analysis, please verify your flow cytometer's capability of exciting and detecting the fluorochrome.
Alexa Fluor® and Pacific Blue™ are trademarks of Life Technologies Corporation.
View full statement regarding label licenses - Excitation Laser
-
Red Laser (633 nm)
- Application Notes
-
NOTE: For flow cytometric staining with this clone, True-Nuclear™ Transcription Factor Buffer Set (Cat. No. 424401) offers improved staining and is highly recommended.
-
Application References
(PubMed link indicates BioLegend citation) - RRID
-
AB_2750492 (BioLegend Cat. No. 126421)
AB_2750493 (BioLegend Cat. No. 126422)
Antigen Details
- Structure
- 50-55 kd protein. Forkhead/winged-helix transcription factor family, contains zinc finger and forkhead domains.
- Distribution
-
Nuclear; expressed in Treg cells.
- Function
- Master regulatory gene in Treg cell development, crucial for immune homeostasis.
- Interaction
- Interacts with DNA
- Cell Type
- Tregs
- Biology Area
- Immunology
- Molecular Family
- Nuclear Markers
- Antigen References
-
1. Ono M, et al. 2007. Nature 446:685.
2. Hori S, et al. 2003. Science 299:1057.
3. Fontenot JD, et al. 2003 Nature Immunol 4:330. - Regulation
- Present at high level in T reg cells. Induced by T cell activation.
- Gene ID
- 20371 View all products for this Gene ID
- UniProt
- View information about FOXP3 on UniProt.org
Related FAQs
- Can I stain whole blood with anti-FOXP3 using your Foxp3 staining kit?
-
It is not recommended. It is best to use PBMCs for this testing.
- Can FOXP3 be costained with cytokines?
-
The larger holes created by the nuclear permeabilization required for FOXP3 may allow cytokines to leak out of the cell, making it harder to detect lowly-expressed cytokines. You may have to use a control where the cells are only permeabilized through the cell membrane.
Other Formats
View All FOXP3 Reagents Request Custom Conjugation| Description | Clone | Applications |
|---|---|---|
| Purified anti-mouse FOXP3 | MF-14 | WB |
| PE anti-mouse FOXP3 | MF-14 | ICFC |
| Alexa Fluor® 488 anti-mouse FOXP3 | MF-14 | ICFC,IHC |
| Alexa Fluor® 647 anti-mouse FOXP3 | MF-14 | ICFC |
| Pacific Blue™ anti-mouse FOXP3 | MF-14 | ICFC |
| Brilliant Violet 421™ anti-mouse FOXP3 | MF-14 | ICFC |
| True-Nuclear™ One Step Staining Mouse Treg Flow™ Kit (FOXP3 Alexa Fluor® 488/CD25 PE/CD4 PerCP) | MF-14 | ICFC |
| Alexa Fluor® 700 anti-mouse FOXP3 | MF-14 | ICFC |
Customers Also Purchased
Compare Data Across All Formats
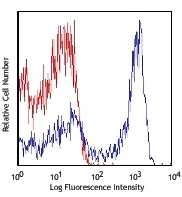
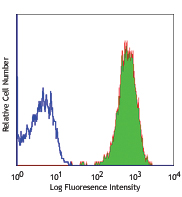
This data display is provided for general comparisons between formats.
Your actual data may vary due to variations in samples, target cells, instruments and their settings, staining conditions, and other factors.
If you need assistance with selecting the best format contact our expert technical support team.
-
Purified anti-mouse FOXP3
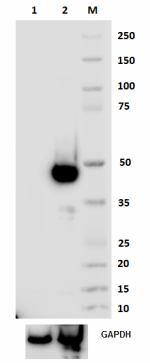
Total cell lysate from 293E (lane 1) and 293E transfected wi... -
PE anti-mouse FOXP3

C57BL/6 splenocytes were surface stained with CD4 APC and th... 
-
Alexa Fluor® 488 anti-mouse FOXP3

C57BL/6 splenocytes were surface stained with CD4 PE and the... 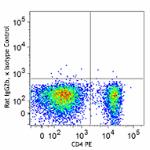
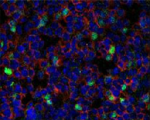
OCT frozen mouse lymph node 5 µm sections were fixed with 4%... -
Alexa Fluor® 647 anti-mouse FOXP3

C57BL/6 splenocytes were surface stained with CD4 PE and the...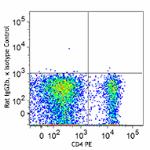 
-
Pacific Blue™ anti-mouse FOXP3
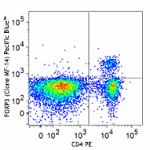
C57BL/6 splenocytes were surface stained with CD4 PE and the...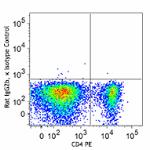 
-
Brilliant Violet 421™ anti-mouse FOXP3
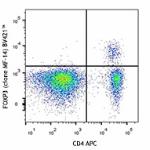
C57BL/6 splenocytes were surface stained with CD4 APC and th... 
-
True-Nuclear™ One Step Staining Mouse Treg Flow™ Kit (FOXP3 Alexa Fluor® 488/CD25 PE/CD4 PerCP)
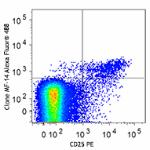
C57BL/6 splenocytes were stained with True-Nuclear™ One Step... 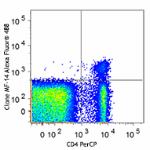
-
Alexa Fluor® 700 anti-mouse FOXP3
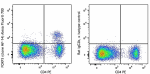

C57BL/6 splenocytes were surface stained with CD4 PE and the...
 Login/Register
Login/Register 












Follow Us