- Clone
- PC10 (See other available formats)
- Regulatory Status
- RUO
- Other Names
- Proliferating Cell Nuclear Antigen, DNA Polymerase δ Auxiliary Protein
- Isotype
- Mouse IgG2a, κ
- Ave. Rating
- Submit a Review
- Product Citations
- publications
| Cat # | Size | Price | Quantity Check Availability | Save | ||
|---|---|---|---|---|---|---|
| 307904 | 100 µg | £155 | ||||
The PC10 monoclonal antibody reacts with proliferating cell nuclear antigen also known as PCNA or the DNA polymerase δ auxiliary protein. PCNA is a 36 kD trimeric ring that acts as a DNA-polymerase sliding clamp expressed in the nucleus of all proliferating cells. A prime function of PCNA appears to be increasing DNA polymerase δ processibility during elongation of the leading strand. PCNA is a useful marker for DNA synthesis and is highly conserved among most species, thus highlighting the very broad reactivity of this antibody.
Product DetailsProduct Details
- Verified Reactivity
- Human, Mouse, Rat
- Antibody Type
- Monoclonal
- Host Species
- Mouse
- Immunogen
- Recombinant rat PCNA
- Formulation
- Phosphate-buffered solution, pH 7.2, containing 0.09% sodium azide.
- Preparation
- The antibody was purified by affinity chromatography, and conjugated with biotin under optimal conditions.
- Concentration
- 0.5 mg/mL
- Storage & Handling
- The antibody solution should be stored undiluted between 2°C and 8°C. Do not freeze.
- Application
-
ICFC - Quality tested
- Recommended Usage
-
Each lot of this antibody is quality control tested by immunofluorescent intracellular staining with flow cytometric analysis. Please follow the Cell Fixation and Permeabilization Protocol Using 70% Ethanol. For flow cytometric staining, the suggested use of this reagent is ≤ 0.125 µg per million cells in 100 µL volume. It is recommended that the reagent be titrated for optimal performance for each application.
- Application Notes
-
Additional reported applications (for the relevant formats) include: immunohistochemical staining2,5,6 of acetone-fixed frozen sections and formalin-fixed paraffin-embedded tissue sections, immunoprecipitation, intracellular flow cytometry3, immunofluorescence microscopy9, and Western blotting10.
-
Application References
(PubMed link indicates BioLegend citation) -
- Ogata K, et al. 1985. J. Immunol. 135:2623.
- Garcia R, et al. 1989. Am. J. Pathol. 134:733.
- Landberg G, et al. 1990. Exp. Cell. Res. 187:111.
- Waseem N, et al. 1990. J. Cell Sci. 96:121.
- Yu C, et al. 1991. Histopathology 19:29.
- Wilkins B, et al. 1992. J. Pathol. 166:45.
- Yang W, et al. 1996. Human Pathol. 27:70.
- Galkowska H, et al. 1996. Vet. Immunol. Immunopathol. 53:329.
- Chou HYE, et al. 2006. J. Biol. Chem. 10:1074.
- Fulvio MD, et al. 2006. Oncogene 25:3932.
- Eswarakumar VP and Schlessinger J. 2007. Proc. Natl. Acad. Sci. USA 104:3937.
- Henkels KM, et al. 2009. Biochem Biophys Res Commun. 389:224. PubMed
- Brobeli A, et al. 2010. Blood Cells Mol Dis. 45:159. PubMed
- Wallace HA, et al. 2014. Development. 141:1332. PubMed
- Mizokami A, et al. 2014. Bone. 69:68. PubMed
- Product Citations
-
- RRID
-
AB_314694 (BioLegend Cat. No. 307904)
Antigen Details
- Structure
- DNA-polymerase sliding clamp, trimeric ring; 36 kD
- Distribution
-
Nuclear, all proliferating cells
- Function
- RAD6-dependent DNA repair pathway; increases DNA polymerase δ processibility during elongation of the leading strand
- Interaction
- PCNA, DNA polymerase δ, Rad6, Rad18, UBC9, MMS2, UBC13, RAD5
- Ligand/Receptor
- Ubiquitination, Sumoylation
- Cell Type
- Neural Stem Cells
- Biology Area
- Cell Biology, Cell Cycle/DNA Replication, DNA Repair/Replication, Immunology, Neuroscience, Neuroscience Cell Markers, Stem Cells
- Molecular Family
- Nuclear Markers
- Antigen References
-
1. Travali S, et al. 1989. J. Biol. Chem. 264:7466.
2. Waseem N, et al. 1990. J. Cell Sci. 96:121.
3. Hall P, et al. 1990. J. Pathol. 162:285.
4. Landberg G, et al. 1991. Cancer Res. 51:4570.
5. Woods A, et al. 1991. Histopathol. 19:21.
6. Hoege C, et al. 2002. Nature 419:135.
7. Yue H, et al. 2003. World J. Gastroenterol. 9:377.
8. Shan B, et al. 2003. J. Biol. Chem. 278:44009. - Gene ID
- 5111 View all products for this Gene ID
- UniProt
- View information about PCNA on UniProt.org
Related Pages & Pathways
Pages
Related FAQs
- How many biotin molecules are per antibody structure?
- We don't routinely measure the number of biotins with our antibody products but the number of biotin molecules range from 3-6 molecules per antibody.
Other Formats
View All PCNA Reagents Request Custom Conjugation| Description | Clone | Applications |
|---|---|---|
| Biotin anti-human/mouse/rat PCNA | PC10 | ICFC |
| PE anti-human/mouse/rat PCNA | PC10 | ICFC |
| Purified anti-human/mouse/rat PCNA | PC10 | ICFC,ICC,IHC,IP,WB |
| Alexa Fluor® 488 anti-human/mouse/rat PCNA | PC10 | ICFC |
| Alexa Fluor® 647 anti-human/mouse/rat PCNA | PC10 | ICFC |
| TotalSeq™-Bn0037 anti-human/mouse/rat PCNA | PC10 | SB |
Compare Data Across All Formats
This data display is provided for general comparisons between formats.
Your actual data may vary due to variations in samples, target cells, instruments and their settings, staining conditions, and other factors.
If you need assistance with selecting the best format contact our expert technical support team.
-
Biotin anti-human/mouse/rat PCNA
-
PE anti-human/mouse/rat PCNA
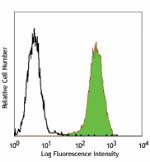
MOLT-4 cells fixed in 70% ethanol then stained with PC10 PE -
Purified anti-human/mouse/rat PCNA
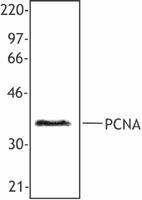

Hela cell nuclear extract was resolved by electrophoresis, t... 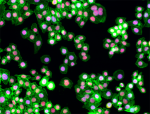
HeLa cells were fixed with 1% paraformaldehyde (PFA) for 10 ... 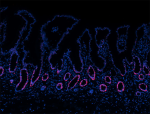
Human paraffin-embedded intestine tissue slices were prepare... -
Alexa Fluor® 488 anti-human/mouse/rat PCNA

MOLT-4 cells fixed in 70% ethanol then stained with PC10 Ale... 
-
Alexa Fluor® 647 anti-human/mouse/rat PCNA
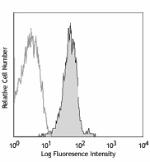
Human peripheral blood lymphocytes fixed with 70% ethanol, t... -
TotalSeq™-Bn0037 anti-human/mouse/rat PCNA
 Login / Register
Login / Register 









Follow Us